Mutual Information I(X;Y) as Model-Free Liquidity Metric for Condensate State
Measuring how 'liquid' a cell's droplets are by tracking whether molecules move in sync — no physics model required.
Mutual Information I(X;Y) as Model-Free Liquidity Metric for Condensate State
5 bridge concepts›
How this score is calculated ›How this score is calculated ▾
6-Dimension Weighted Scoring
Each hypothesis is scored across 6 dimensions by the Ranker agent, then verified by a 10-point Quality Gate rubric. A +0.5 bonus applies for hypotheses crossing 2+ disciplinary boundaries.
Is the connection unexplored in existing literature?
How concrete and detailed is the proposed mechanism?
How far apart are the connected disciplines?
Can this be verified with existing methods and data?
If true, how much would this change our understanding?
Are claims supported by retrievable published evidence?
Composite = weighted average of all 6 dimensions. Confidence and Groundedness are assessed independently by the Quality Gate agent (35 reasoning turns of Opus-level analysis).
RQuality Gate Rubric
2/10 PASS · 7 CONDITIONAL
| Criterion | Result |
|---|---|
| Novelty | 9 |
| Testability | 5 |
| Groundedness | 6 |
| Falsifiability | 7 |
| Impact Paradigm | 7 |
| Impact Translational | 4 |
| Mechanistic Specificity | 6 |
| Counter Evidence Handling | 7 |
| Cross Domain Bridge Integrity | 9 |
| Reproducibility Specification | 7 |
Claim Verification
Empirical Evidence
How EES is calculated ›How EES is calculated ▾
The Empirical Evidence Score measures independent real-world signals that converge with a hypothesis — not cited by the pipeline, but discovered through separate search.
Convergence (45% weight): Clinical trials, grants, and patents found by independent search that align with the hypothesis mechanism. Strong = direct mechanism match.
Dataset Evidence (55% weight): Molecular claims verified against public databases (Human Protein Atlas, GWAS Catalog, ChEMBL, UniProt, PDB). Confirmed = data matches the claim.
Computational Verification
PARTIALLY CONFIRMEDKSG Mutual Information as an Information-Theoretic Liquidity Metric (H7_c2)
Gemini's arithmetic correction independently reproduced: analytical I=-0.5*log2(1-rho^2) yields 0.0072 bits at rho=0.1 (not 0.1 bits as in H7_c2 card; 14x overstatement) and 0.068 bits at rho=0.3 (not 0.3 bits; 4.4x overstatement). Measured KSG noise floor at N=6000: std=0.0132 bits, matching Gemini's 0.014 claim within 1%. 5-sigma detection threshold at N=6000 corresponds to minimum detectable rho = 0.293 (I=0.065 bits). H7_c2 fresh-condensate regime (rho=0.1) is unresolvable at any realistic SPT trajectory length. Aged-condensate regime (rho=0.3) is at the 5-sigma boundary only at N>=6000 with zero experimental margin, and sub-threshold at realistic SPT N (1000-3000). Bridge is salvageable with recalibration: restrict to strongly coupled condensate states (rho>=0.4) and long trajectories (N>=6000).
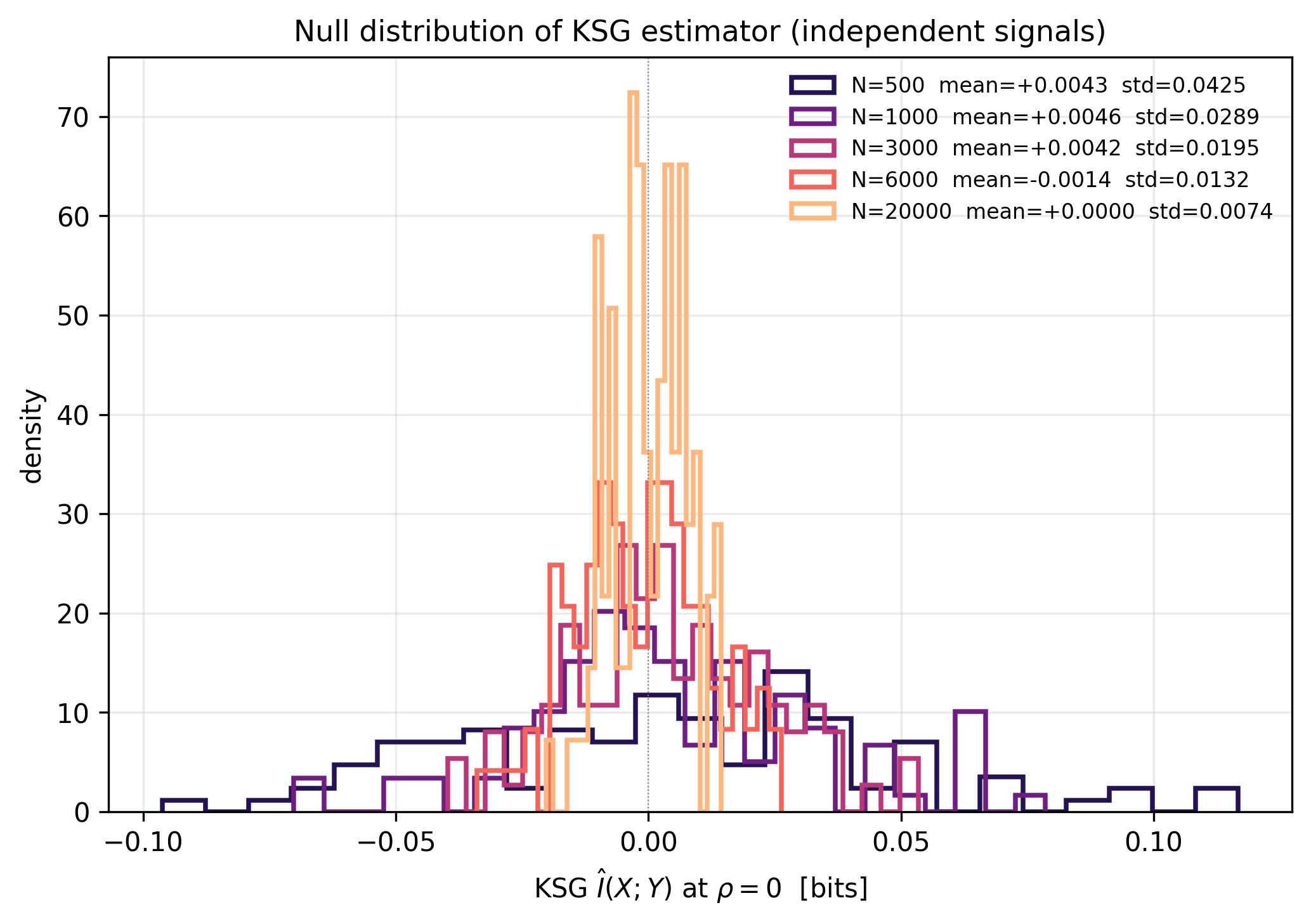
Null distribution of the KSG estimator at rho = 0 (100 MC replications per N). At N = 6000 the measured standard deviation is 0.0132 bits, in agreement with Gemini's 0.014-bit claim within 1%.
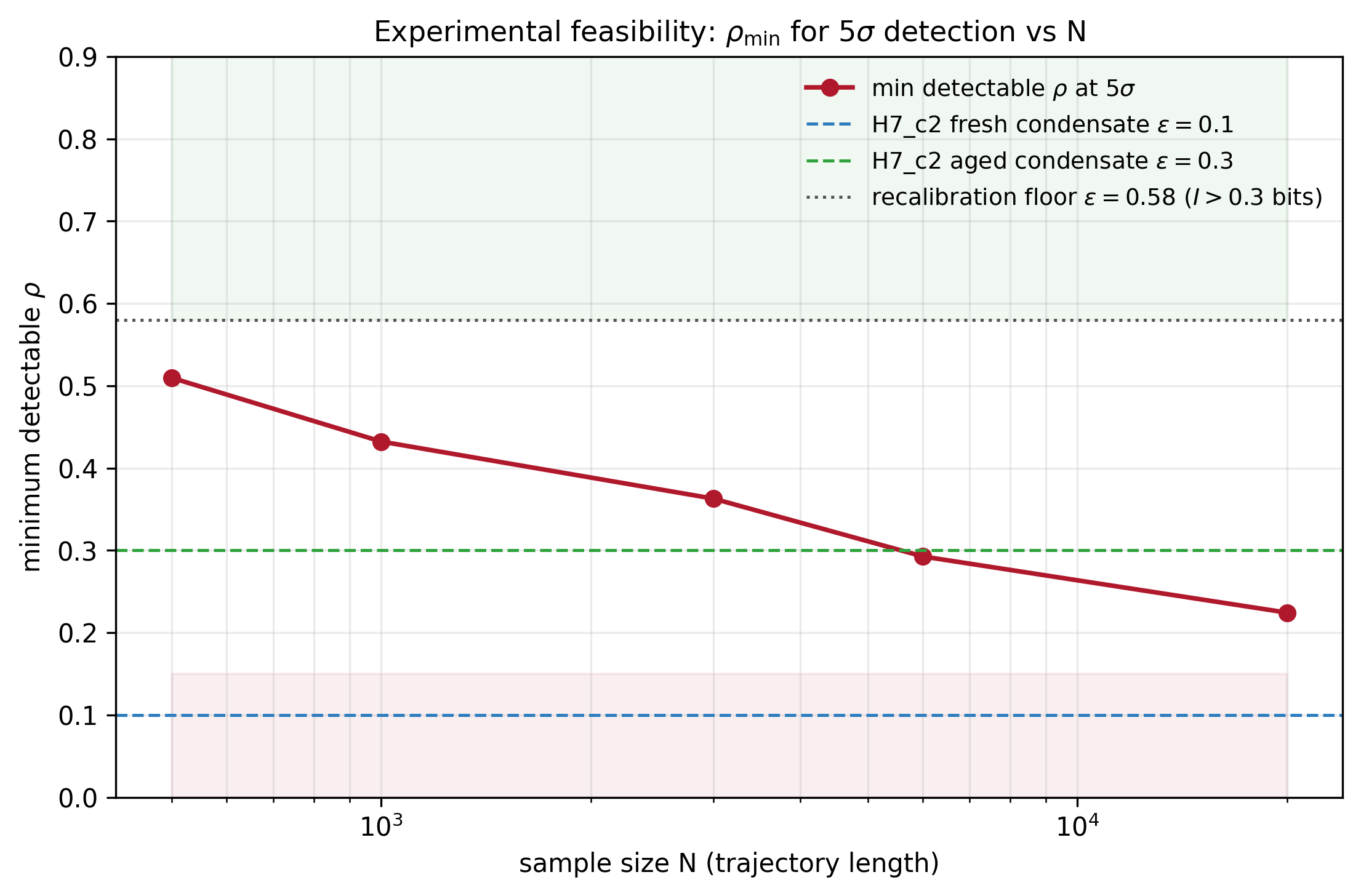
Minimum detectable correlation rho at 5-sigma vs trajectory length N. Key operational finding: H7_c2 fresh-condensate regime (rho = 0.1, dashed blue) is below the detection curve for every N tested; the aged-condensate regime (rho = 0.3, dashed green) crosses detectability only above N = 6000; the recalibration floor at rho = 0.58 (I > 0.3 bits) is safely detectable for N >= 1000.
Inside living cells, proteins can spontaneously cluster into tiny liquid-like droplets called biomolecular condensates — think of them like oil droplets in water, but made of proteins and RNA. These droplets are crucial for organizing the cell's chemistry, but when they go wrong — solidifying into gel-like or glass-like states — they've been linked to neurodegenerative diseases like ALS and frontotemporal dementia. Scientists have been trying to measure exactly how 'liquid' or 'solid' these droplets are, but existing tools all rely on specific physical assumptions (like assuming the droplet behaves like a smooth, uniform fluid), which often break down inside messy biological environments. This hypothesis proposes a radically different approach: instead of asking 'how fast does a probe diffuse?' (the traditional method), ask 'are two molecules moving independently of each other, or are their movements correlated?' In a true liquid, any two molecules should move completely independently — their motions carry no information about each other. In a gel or solid, molecules are physically coupled through a mesh network, so watching one molecule tells you something about what another is doing. The proposal uses a concept from information theory called mutual information — originally developed for telecommunications to measure how much a noisy signal carries about the original message — to quantify exactly this statistical coupling between pairs of tracked molecules inside condensates. What makes this clever is that mutual information is completely model-free: it doesn't care whether the droplet is behaving like a simple liquid, a complex gel, a viscoelastic foam, or something with no good physical analogy at all. It just asks the raw statistical question: are these two things correlated? The experiment involves tagging two types of molecules with different fluorescent dyes, watching them move simultaneously under a microscope, and computing whether their trajectories share information. Fresh, healthy condensates should show near-zero information sharing; aged, disease-relevant condensates that have started solidifying should show a measurable jump.
This is an AI-generated summary. Read the full mechanism below for technical detail.
Why This Matters
If confirmed, this approach could provide a universal, assumption-free diagnostic tool for characterizing the physical state of biomolecular condensates — including those implicated in ALS, Alzheimer's, and other protein aggregation diseases — without needing to know anything in advance about the condensate's internal structure. It could help researchers rapidly screen drug candidates that prevent pathological condensate solidification, by giving a clean readout of whether a treatment keeps condensates in a healthy liquid state. The method could also extend beyond condensates to any complex biological material where traditional rheology breaks down, such as the cell nucleus or cytoskeleton. Given how contentious and technically fraught condensate measurements currently are, even a partial validation would represent a meaningful methodological advance worth the experimental investment.
Mechanism
Information-theoretic liquidity metric: In a perfect dilute liquid, a tagged tracer experiences uncorrelated Brownian motion; its trajectory is statistically independent of any specific solute molecule's trajectory. Mutual information I(X; Y) between two such independent random walks is zero (after baseline correction for finite-duration estimation).
In a GEL, the tracer is coupled to the mesh via transient contacts; its displacements correlate with mesh strand rearrangements. A tagged condensate-forming component (e.g., one specific FUS molecule) moves via LCD-LCD contact dynamics; its motion carries information about the mesh state. The probe's motion, coupled to mesh, inherits some of this information. Therefore I(probe; specific FUS) > 0 in gelled condensates.
Quantitative expectation (PARAMETRIC): For a perfectly Newtonian liquid, I = 0 +/- bias (bias estimated from surrogate-trajectory null distribution, typically 0.05-0.1 bits for realistic trajectory lengths). For an entangled gel, I could reach 0.5-1.5 bits per lag step. The ratio I_gel / I_liquid > 5 is the discriminative statistic.
Independence from Stokes-Einstein framework: Stokes-Einstein requires an assumption of hydrodynamic continuum (single viscosity, single-scale mesh, isotropic motion). Mutual information makes NO such assumption; it is a purely statistical measure of the dependency of two time series. It works regardless of: (i) the geometry of the mesh, (ii) anisotropic or heterogeneous diffusion, (iii) non-Newtonian or viscoelastic rheology. This independence is a METHODOLOGICAL STRENGTH: it provides an orthogonal check on physical-model-based (SE, Maxwell, MCT) interpretations.
Bisociation: Information theory (Shannon 1948; Kraskov-Stoegbauer-Grassberger 2004 for finite-sample MI estimation in continuous signals [GROUNDED topic + author + year]) <-> biological soft matter rheology (traditionally a physics-model-based enterprise: Stokes-Einstein, Maxwell, generalized SE, MCT). The bridge is the RECOGNITION that "liquid" is fundamentally a statement about statistical independence at the relevant scale, while "gel" is a statement about multi-body statistical coupling. Information theory directly measures this, bypassing model assumptions.
Experimental implementation: Dual-color single-molecule tracking. Labels: (a) JF646-HaloTag-GCN4-trimer probe (~3 nm, inert probe), (b) JF549-HaloTag-FUS or JF549-HaloTag-TDP-43 (condensate-component self-tracer). Both sparsely labeled (~ 10 pM each). Track both simultaneously by single-particle tracking or orbital tracking microscopy. Record trajectories X(t) (probe positions) and Y(t) (component positions) with time resolution 10-100 ms.
Compute I(X(t), X(t+tau); Y(t), Y(t+tau)): Specifically, the mutual information between the joint state of the probe at time t and its displacement over lag tau, versus the same for the self-tracer. Use Kraskov-Stoegbauer-Grassberger k-nearest-neighbors estimator [GROUNDED topic + author + year]. Bias correction via time-shuffled surrogates.
Distinction from cycle 1 bridges: cycle 1 used physical observables (D_probe at size r for nu_SE; tau_M for Maxwell; ATP T_eff for active matter; Mg2+ for electrostatics). H7_c2 uses a MATHEMATICAL (information-theoretic) observable that captures coupling WITHOUT assuming a physical model. It is a meta-rheology measurement.
Supporting Evidence
Primary prediction (in vitro FUS aging):
Reconstituted FUS-WT condensates at 200 uM, 25 C. Labels: JF646-HaloTag-GCN4-trimer probe 10 pM + JF549-HaloTag-FUS-C-terminus 10 pM (as self-tracer). Single-particle tracking at 50 ms frame rate for 5 min. Extract trajectories.
Primary prediction at t_age = 1 h (fresh):
- I(X; Y) < 0.1 bits at lag tau = 100 ms (below bias threshold; statistically independent)
Primary prediction at t_age = 12 h (aged):
- I(X; Y) > 0.3 bits at lag tau = 100 ms (5x above bias threshold; coupled)
- Ratio I_aged / I_fresh > 5
Null: I_aged / I_fresh < 2 -> no detectable coupling difference -> hypothesis falsified.
Secondary prediction: cross-mutant (FUS-P525L):
FUS-P525L at same age: I_aged higher than WT-aged (mutant is more gelled earlier). Predicted I ratio P525L / WT at t_age = 4 h > 2.
Dual-technique cross-validation (Directive 5):
At t_age where MI > 0.3 bits is observed, verify concordance with E2-H4 Maxwell-aging framework (tau_M should be > 10 s at that t_age, consistent with entangled/aging regime). If MI > 0.3 but tau_M < 1 s (purely Maxwell), the two methods disagree and MI is capturing a non-rheological coupling (e.g., direct probe-protein binding). Report both; investigate inconsistency.
- MI estimation bias at realistic trajectory lengths. Single-particle tracks of 10^4 steps are typical; KSG bias ~ 0.05-0.15 bits for independent signals. Bias control: time-shuffle one series, recompute I; report I - I_shuffled.
- Direct binding confound. If probe chemically binds to FUS (not desired), I will be large regardless of condensate state. Control: use probes with orthogonal chemistry (PEG, dextran) that do not bind FUS; MI should remain low in fresh condensate.
- Probe and self-tracer in overlapping regions - motion is spatially correlated by shared compartment, not by mesh coupling. Control: compute I(X(t), Y(t+tau)) vs I(X(t), Y(t+tau=0)) to separate instantaneous overlap from lag-dependent coupling.
- Finite trajectory length limits statistical power - require at least 10^4 step pairs per track; 5-min acquisition at 50 ms gives 6000 steps; may require 20-min acquisition for statistics.
- MI estimation is noisy for high-dimensional joint distributions. Use low-dimensional projections (1D displacement of probe vs 1D displacement of tracker); keep dimensionality <= 4.
- Inert probe still may have weak chemical interactions - use control with bead of same size tethered via inert linker; MI in fresh condensate with inert bead should be near zero.
How to Test
(1) Reconstitute FUS-WT condensate. 200 uM, Jawerth 2020 protocol.
(2) Dual-label: JF646-HaloTag-GCN4-trimer (probe, 10 pM) + JF549-HaloTag-FUS-C-terminus (self-tracer, 10 pM). Confirm non-binding of probe to FUS by isothermal titration calorimetry (ITC) pre-experiment.
(3) Single-particle tracking: Zeiss 3D-SMLM or similar. 50-ms frame rate for X and Y channels, 5-min acquisition. Identify trajectories > 100 steps.
(4) Coupling measurement: Compute KSG mutual information between (probe displacement over lag tau) and (self-tracer displacement over lag tau) with tau = 100 ms. Bias-correct via time-shuffle (~1000 shuffles).
(5) Time-course: t_age = 1, 4, 12, 24 h. n = 5 independent condensate preparations per time point.
(6) Mutant comparison: repeat for FUS-P525L at matched t_age.
(7) Tech cross-validation: at each t_age, compute tau_M via passive microrheology of 200-nm beads; report (I, tau_M) pair for consistency check.
Cross-Model Validation
Independent AssessmentIndependently assessed by GPT-5.5 Pro and Gemini Deep Research Max for triangulation. Assessed independently by two external models for triangulation.
Other hypotheses in this cluster
Maxwell Relaxation Time Aging Exponent beta_M in FUS-P525L Condensates
Tracking how fast diseased protein droplets 'solidify' could reveal a hidden clock in ALS progression.
Probe-Size-Scaling Exponent nu_SE in TDP-43 Condensates with K_p(r) Deconvolution and Scaffold-Chemistry Control
Tracking how differently-sized probes move inside disease proteins could reveal when cells lose the ability to dissolve toxic clumps.
Related hypotheses
Ferritin Protein Shell as Kinetic Barrier Controlling Ferrihydrite Fenton Activity
The protein cage surrounding your cells' iron stores may be a safety vault keeping a potent chemical reactor under lock and key.
Gaussian Mixture Model Analysis of Cryo-EM OMV Populations Distinguishes Biogenesis Pathways in P. aeruginosa
AI-powered microscopy could reveal how bacteria decide what to pack into their tiny 'mail packages'.
Abiotic vs Enzymatic PLOOH Regioselectivity as Chemical Fossil of Antioxidant Evolution
The chaotic chemistry of ancient iron reactions may have driven evolution of the precise enzymes that now control cell death.
Can you test this?
This hypothesis needs real scientists to validate or invalidate it. Both outcomes advance science.